Quantitative Estimation of Recurrent Genomic Alterations
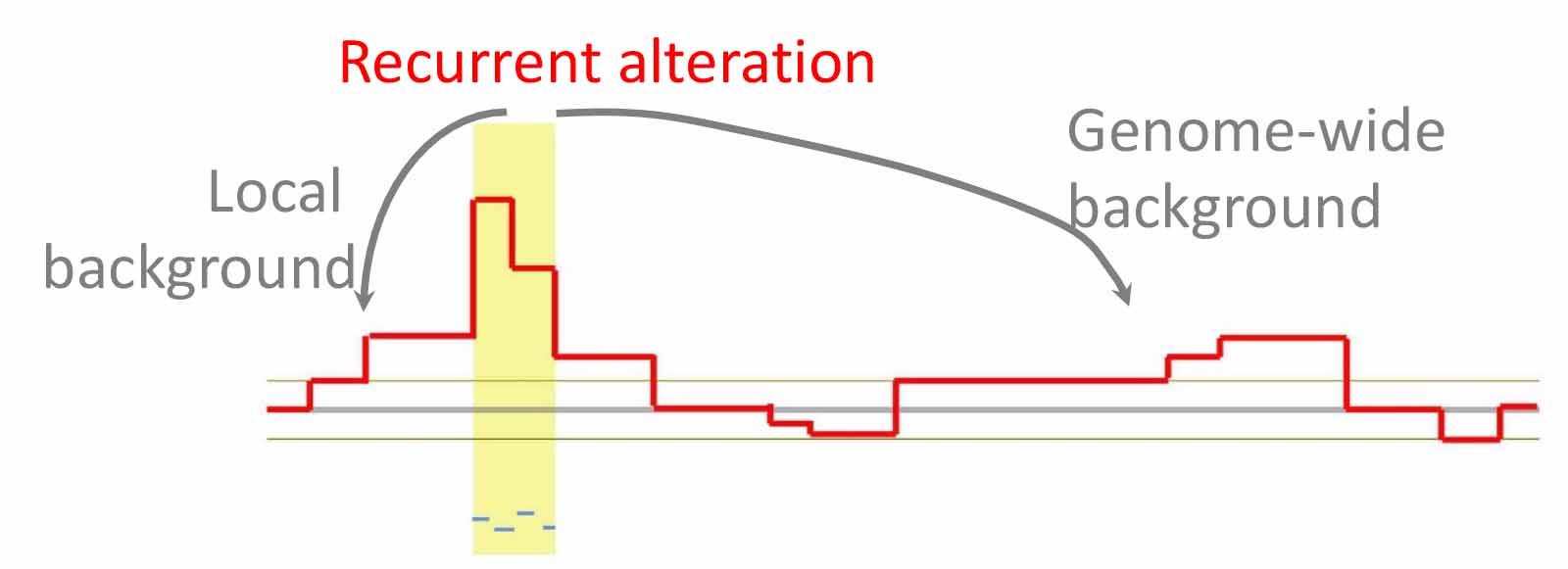
Tumorigenesis is driven by the acquisition of somatic alterations in the genome, including copy number alterations, single nucleotide substitutions, small insertions/deletions, and structural rearrangements. The recurrence of genomic alterations in independent cancer patients is a strong indicator of their functional impact, as the frequency of an alteration statistically higher than the background suggests the presence of selective pressures during tumor cell evolution.
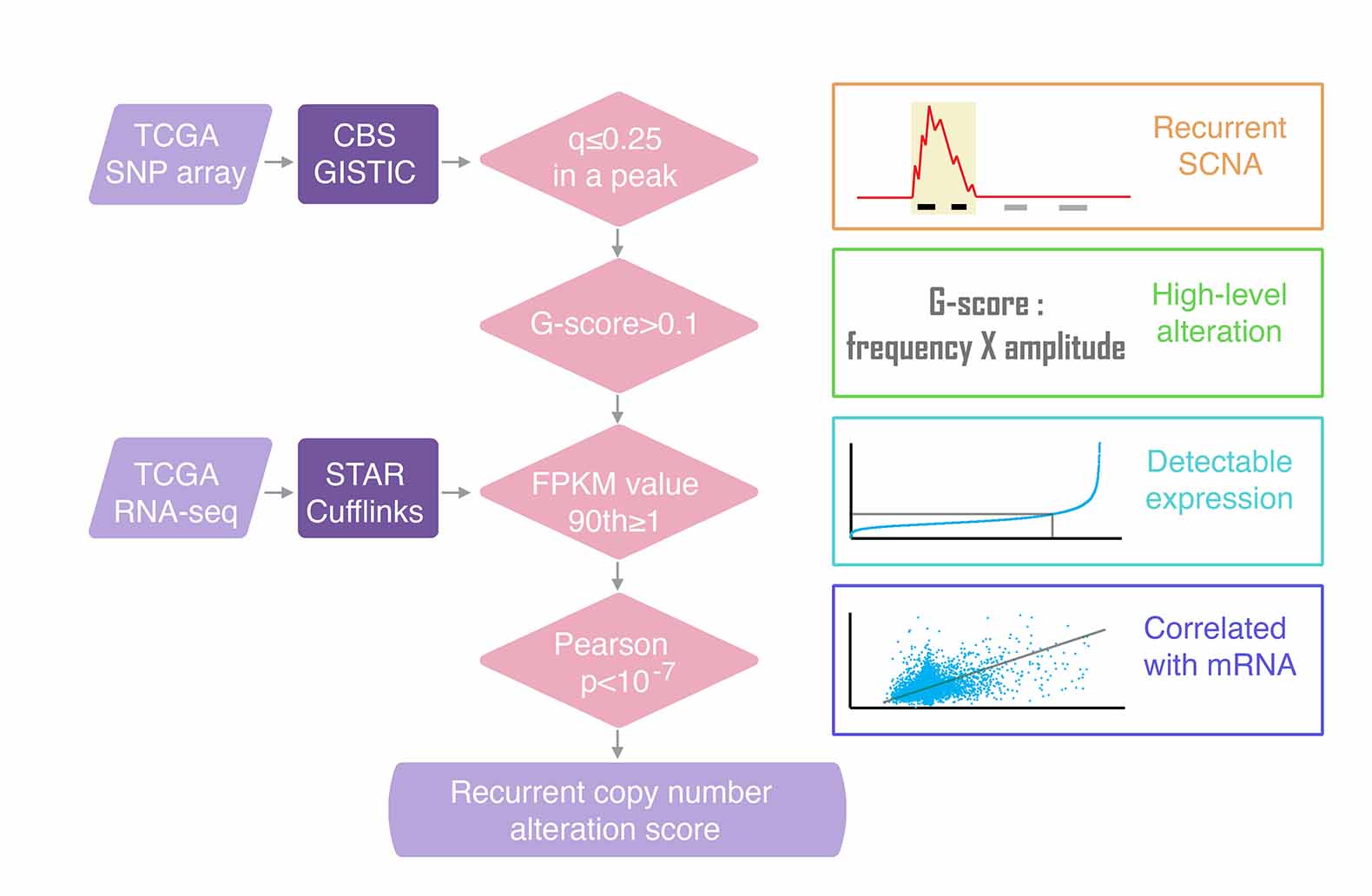
Somatic copy number alterations
To characterize somatic copy number alterations (SCNAs), we analyzed single nucleotide polymorphism array profiles from TCGA. SCNAs in cancer genomes are classified into two types: focal SCNAs, confined to small genomic regions, and broad (arm-level) SCNAs, which encompass large fragments or whole chromosomal arms. Recurrent focal SCNAs, containing only a few genes, often exhibit high-amplitude variation and occur more frequently than the background, indicating additional selective pressures during tumor cell evolution. This suggests that the genes in these regions likely play causal roles in tumorigenesis.
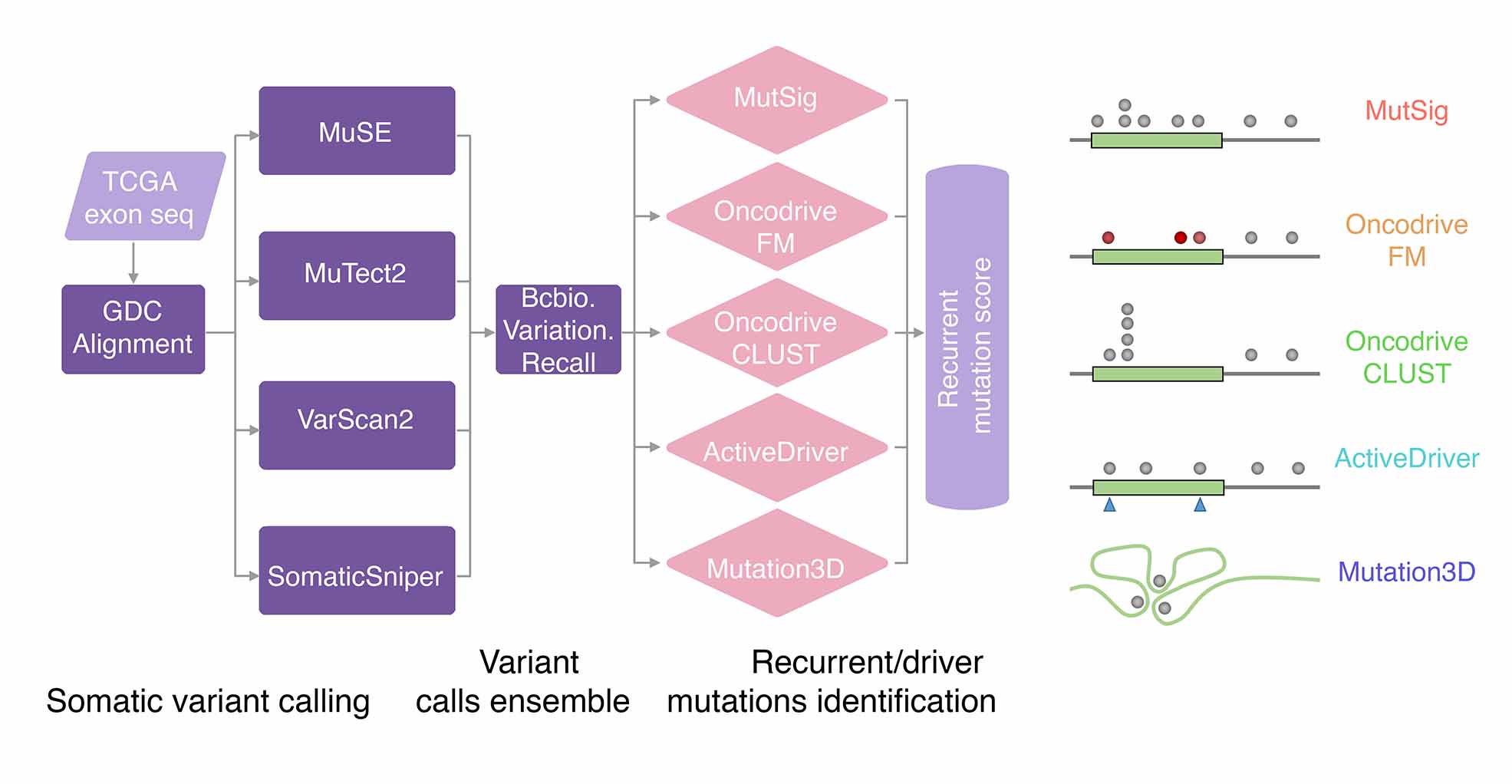
Somatic mutations
For somatic mutations (single nucleotide variants and indels), we analyzed whole exome sequencing (WES) profiles from TCGA. Despite numerous methods developed to enhance mutation calling accuracy, somatic variant calling in tumor specimens remains challenging due to sample heterogeneity and copy number alterations. To improve sensitivity and precision, we utilized the mutation call set generated via an ensemble calling strategy by the MC3 (Multi-Center Mutation Calling in Multiple Cancers) project. This involved combining variant call format (VCF) mutation profiles generated independently by seven different methods (MuTect, Indelocator, Pindel, MuSE, Radia, Varscan2, and SomaticSniper) into a single call set, followed by the application of automated filters. Recurrently mutated genes, exhibiting significantly higher mutation rates than the background, indicate positive selection during tumorigenesis. We integrated five complementary methods to predict recurrent somatic mutations reliably. The mutation index, ranging from 0 to 5, was defined as the number of methods with a positive result for a given gene in a certain cancer type. Additionally, a mutation score (M-score), considering both the mutation index and the mutation frequency across samples, was estimated for each gene. Genes identified by at least two methods (mutation index ≥2) were considered recurrently mutated.
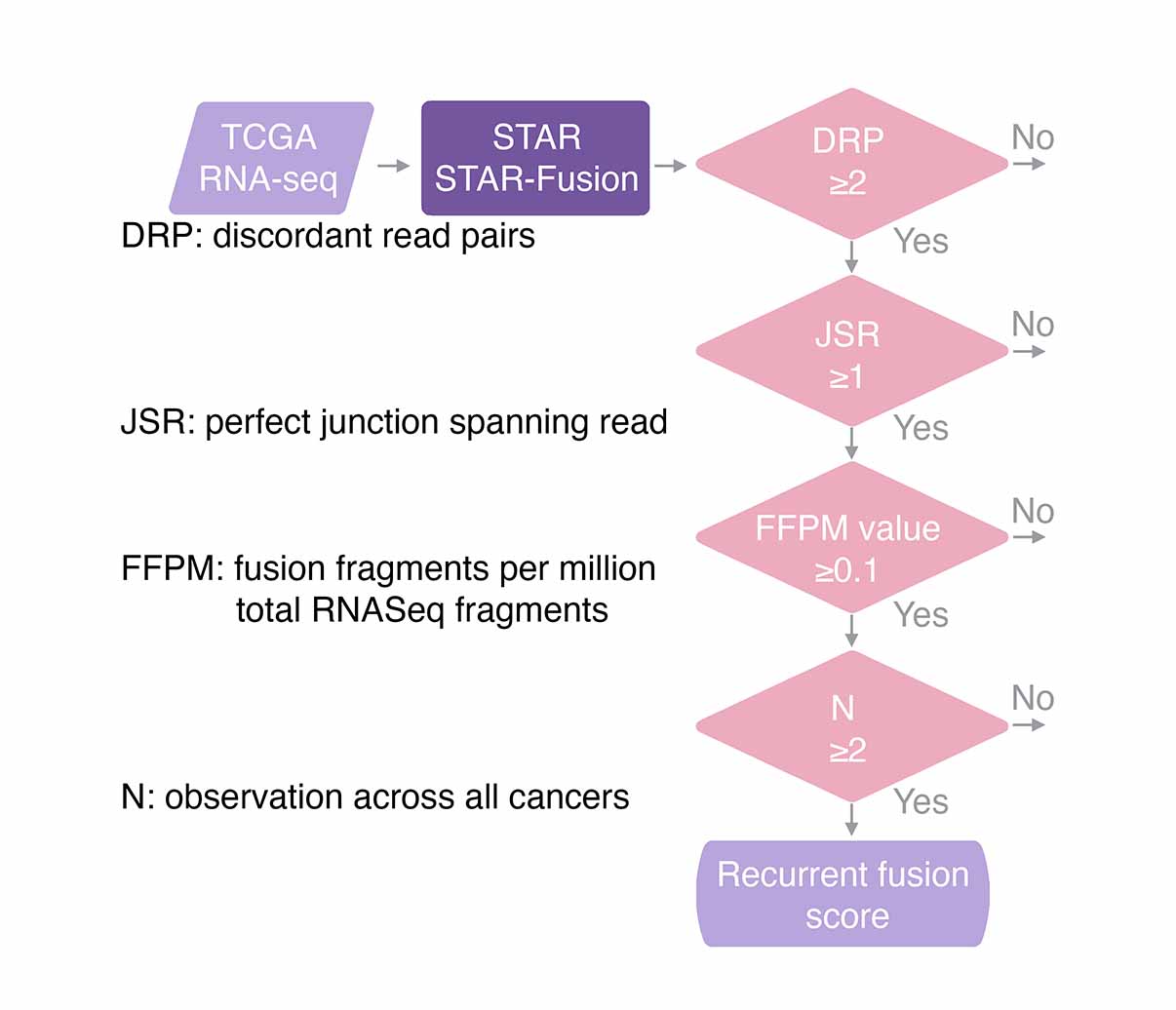
Transcript fusions
To characterize transcript fusions, we retrieved gene fusion data from TCGA using the TumorFusions database.
Detailed methods can be found in our previous publications:
1. Jiang J, Yuan J, Hu Z, Zhang Y, Zhang T, Xu M, Long M, Fan Y, Tanyi JL, Montone KT, Tavana O, Vonderheide RH, Chan HM, Hu X, Zhang L. Systematic illumination of druggable genes in cancer genomes. Cell Rep. 2022 Feb 22;38(8):110400. doi: 10.1016/j.celrep.2022.110400. PubMed PMID: 35196490; PubMed Central PMCID: PMC8919705.
2. Hu Z, Yuan J, Long M, Jiang J, Zhang Y, Zhang T, Xu M, Fan Y, Tanyi JL, Montone KT, Tavana O, Chan HM, Hu X, Vonderheide RH, Zhang L. The Cancer Surfaceome Atlas integrates genomic, functional and drug response data to identify actionable targets. Nat Cancer. 2021 Dec;2(12):1406-1422. doi: 10.1038/s43018-021-00282-w. Epub 2021 Dec 13. PubMed PMID: 35121907; PubMed Central PMCID: PMC9940627.
3. Hu Z, Zhou J, Jiang J, Yuan J, Zhang Y, Wei X, Loo N, Wang Y, Pan Y, Zhang T, Zhong X, Long M, Montone KT, Tanyi JL, Fan Y, Wang TL, Shih IM, Hu X, Zhang L. Genomic characterization of genes encoding histone acetylation modulator proteins identifies therapeutic targets for cancer treatment. Nat Commun. 2019 Feb 13;10(1):733. doi: 10.1038/s41467-019-08554-x. PubMed PMID: 30760718; PubMed Central PMCID: PMC6374416.
