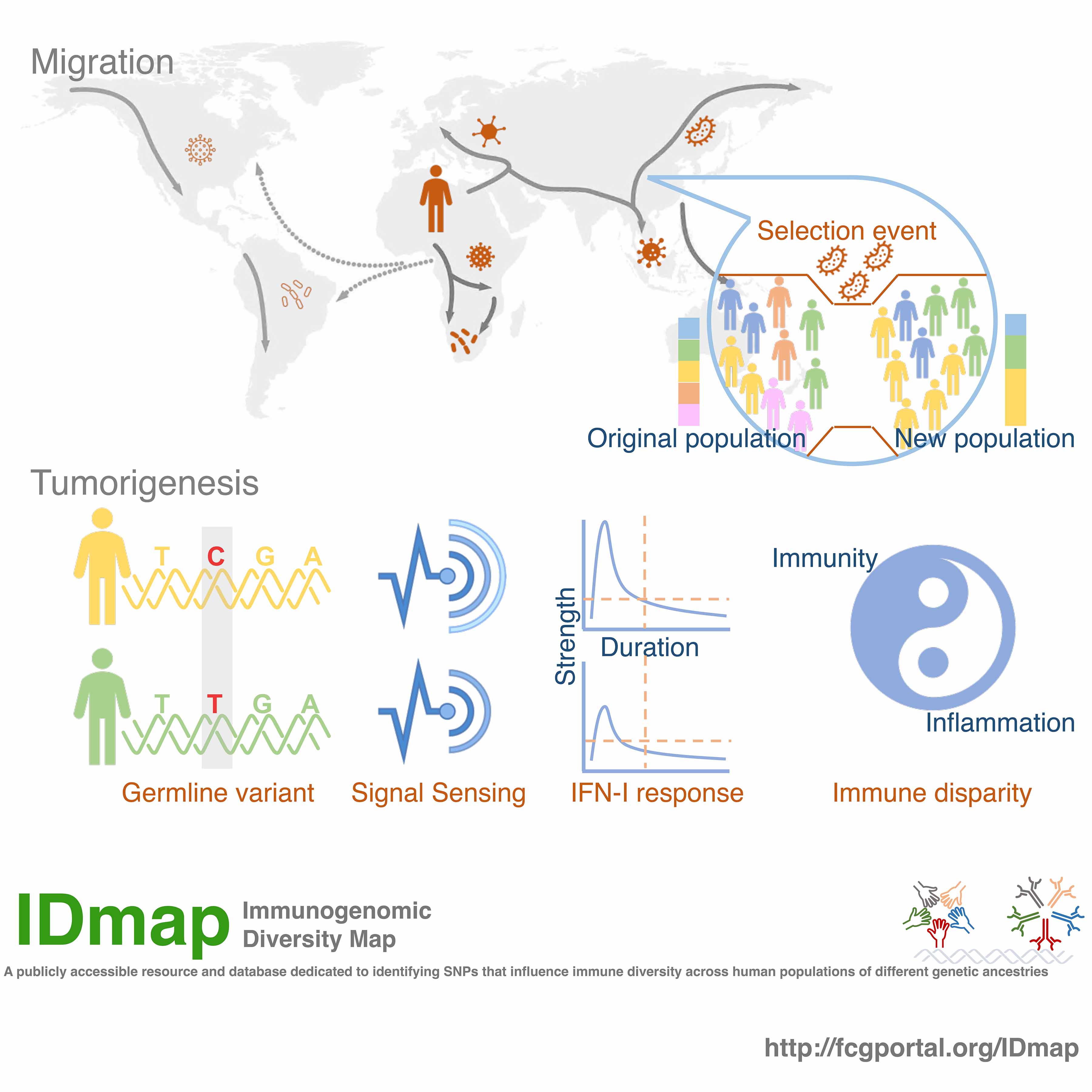
As human populations migrate to diverse geographical regions, they encounter a broad spectrum of pathogens, leading to pronounced natural selection pressures on the immune system. Through the analysis of nonsynonymous single nucleotide polymorphisms (nsSNPs) in human genomes across major geographically structured populations, we observed increased nsSNP variation in immune-related genes, indicative of evolutionary adaptations in response to historical pathogen-driven selection. In contrast, cancer-related genes display lower nsSNP variation, suggesting evolutionary conservation and their critical roles in cellular processes. By investigating a large collection of cancer cell lines, we identified 24 nsSNPs within genes encoding pattern recognition receptors associated with distinct type I interferon (IFN-I) activity.
Among these, nsSNPs in STING1 exhibit the highest impact score, reflecting their
significant variation across human populations and a substantial influence on intrinsic IFN-I activity in cancer cells. Subsequent experimental and computational analyses demonstrated functional impacts of STING1 variants on the intrinsic cGAS-STING1-IFN signaling in cancer cells and their potential influence on tumor immunity in cancer patients. Finally, we conducted a genome-wide characterization of immunogenomic diversity genes across genetic ancestry populations and established a publicly accessible database, the Immunogenomic Diversity Map (IDmap). Our study indicates that alongside socioeconomic, cultural, and environmental determinants, genetic ancestry-related germline variances may substantially influence cancer immunity and the response to immune therapy treatments.